三甲病院 リアル外来処方箋
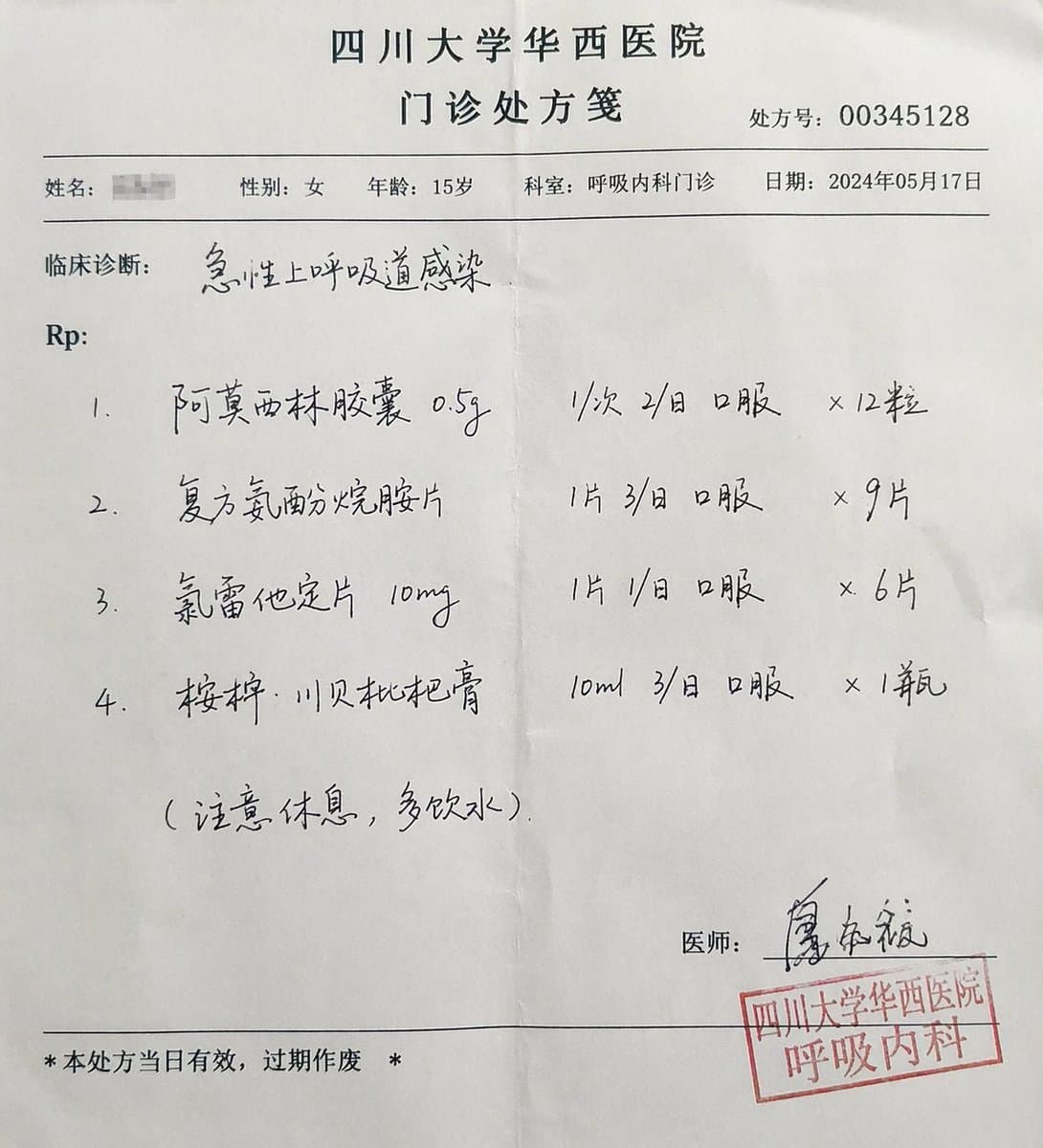
他のインフォグラフィック
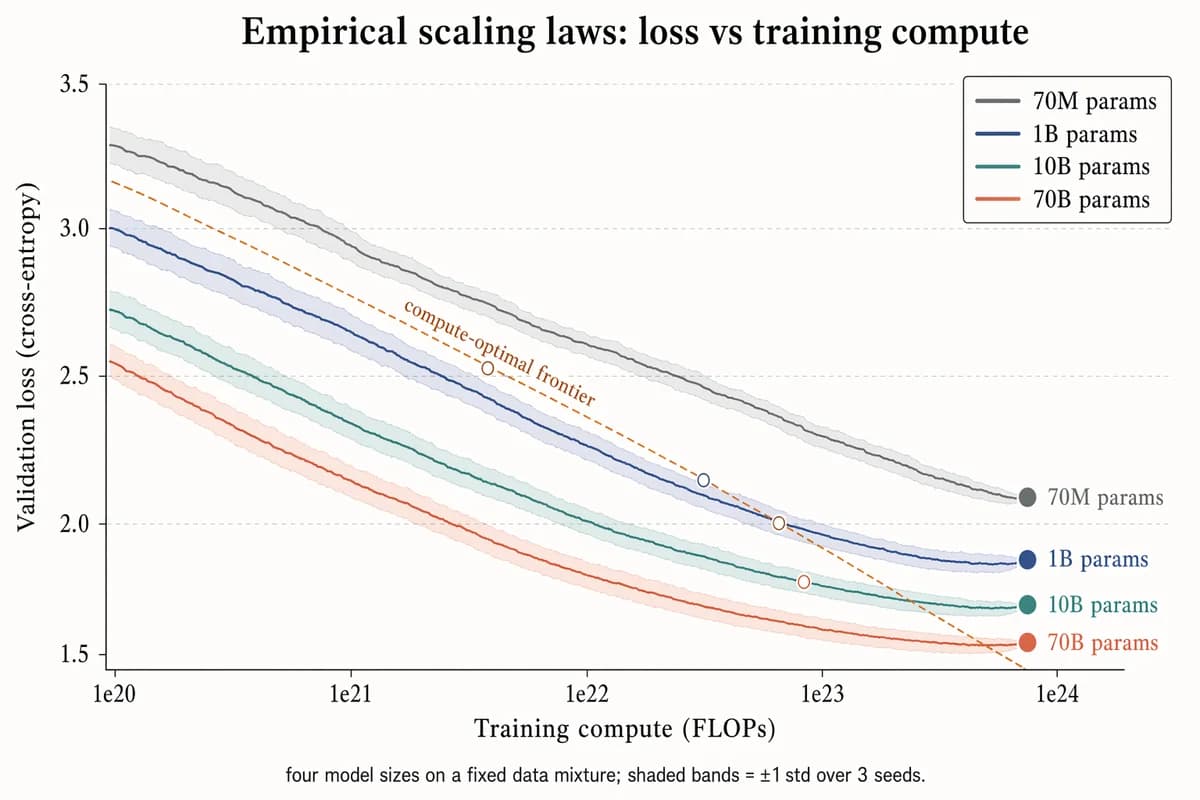
経験的スケーリング則 プロット
Landscape 16:9 log-scaled plot of training loss vs compute, four curves for different model sizes. X-axis "Training compute (FLOPs)" with log ticks "1e20", "1e21", "1e22", "1e23", "1e24". Y-axis "Validation loss (cross-entropy)" with linear decreasing ticks "3.5", "3.0", "2.5", "2.0", "1.5". Four descending curves with ±1σ shaded bands, labels near tails: "70M params" (slate gray), "1B params" (muted navy), "10B params" (dusty teal), "70B params" (soft terracotta). Warm-copper dashed diagonal line labeled "compute-optimal frontier"; open circles at isoflop crossover points. Legend box top-right. Title: "Empirical scaling laws: loss vs training compute". Subtitle: "four model sizes on a fixed data mixture; shaded bands = ±1 std over 3 seeds."
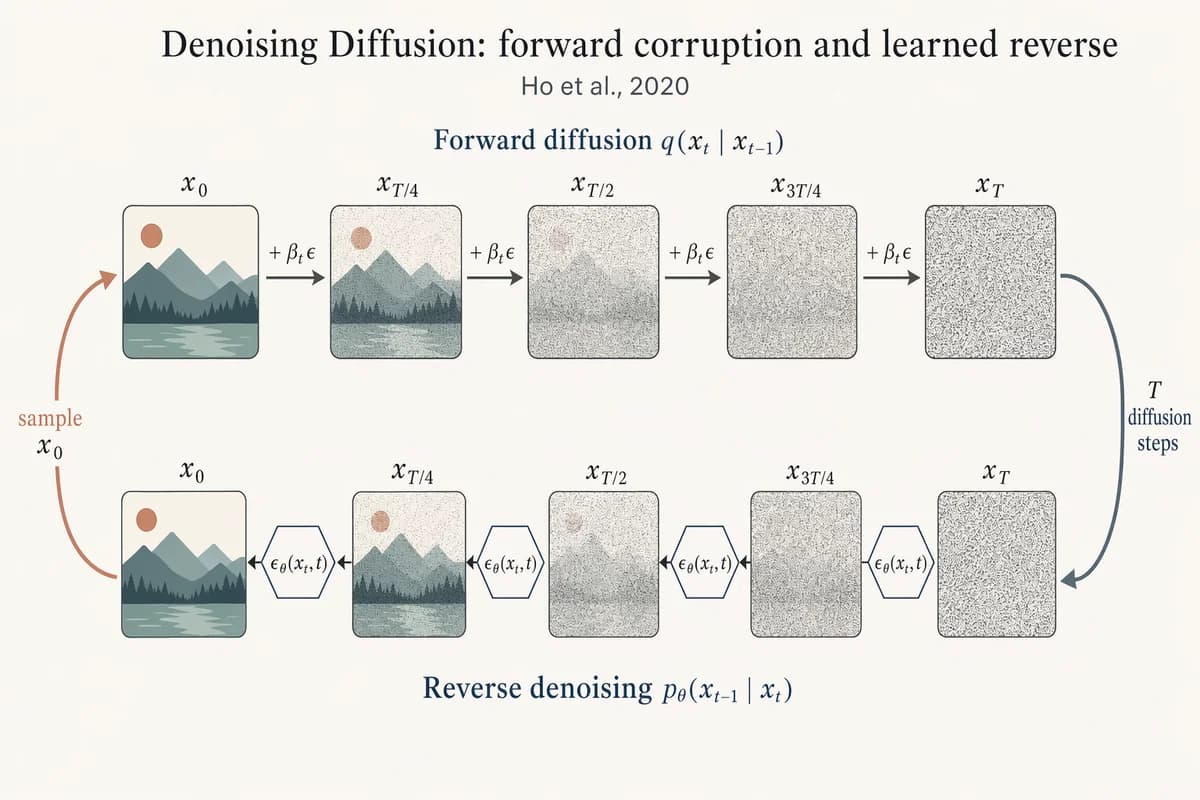
デノイジング拡散モデル 順方向/逆方向チェーン
Landscape 16:9 academic figure of diffusion forward + reverse chains, two horizontal chains stacked vertically. TOP chain (left→right) labeled "Forward diffusion q(x_t | x_{t-1})": five frames "x_0", "x_{T/4}", "x_{T/2}", "x_{3T/4}", "x_T" progressing from a crisp small mountain-sun landscape to pure Gaussian noise. Arrows between frames labeled "+ β_t ε". BOTTOM chain (right→left) labeled "Reverse denoising p_θ(x_{t-1} | x_t)": same five frames in reverse, with a small hexagonal ε_θ(x_t, t) block between each pair. Far-right curved arrow "T diffusion steps" connecting top-right to bottom-right; far-left curved arrow "sample x_0" connecting bottom-left to top-left. Title: "Denoising Diffusion: forward corruption and learned reverse". Subtitle: "Ho et al., 2020".
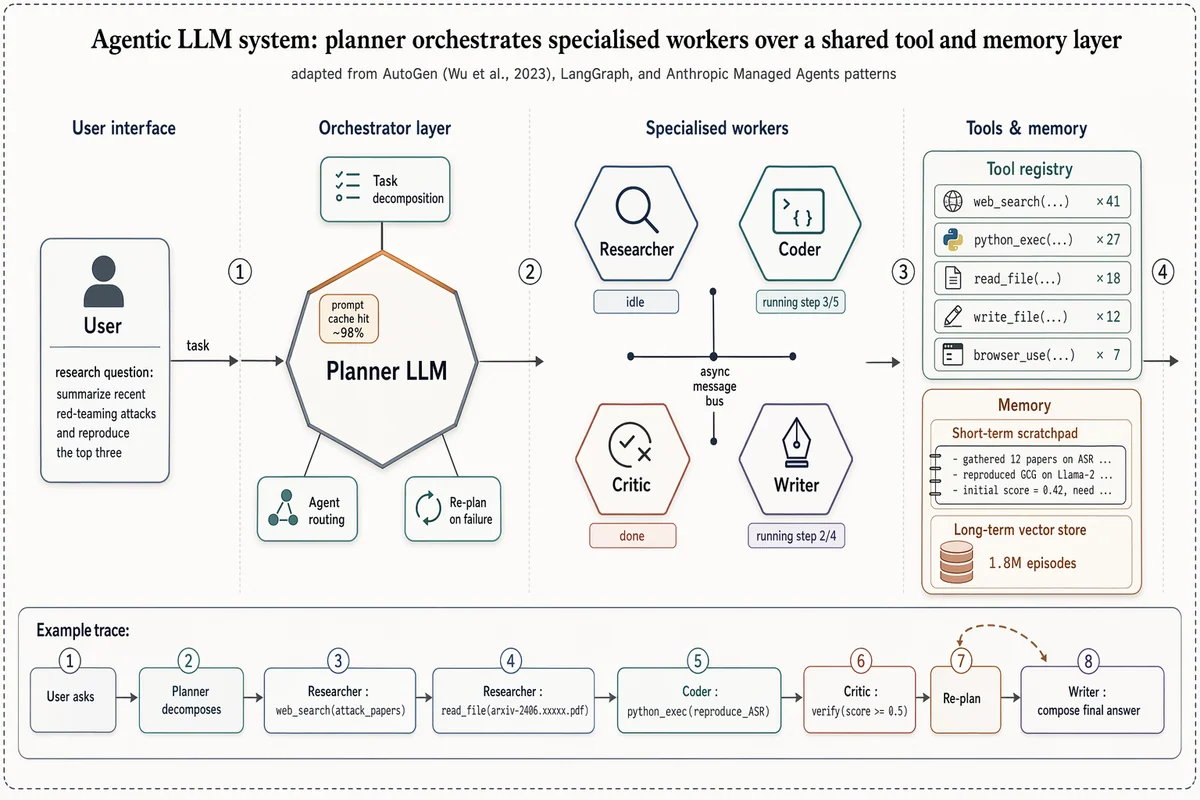
マルチエージェントLLM システムアーキテクチャ
Landscape 16:9 high-fidelity systems figure of a multi-agent LLM architecture, in the style of a richly detailed AutoGen / LangGraph / Anthropic Managed Agents Figure 1. Subtle drop-shadows, warm-copper highlights, numbered flow markers ①②③④. ZONE 1 — "User interface": rounded user box with placeholder task "research question: summarize recent red-teaming attacks and reproduce the top three". ZONE 2 — "Orchestrator layer": central hexagonal hub "Planner LLM" with warm-copper top edge. Three satellite chips: "Task decomposition", "Agent routing", "Re-plan on failure". Small inset chip "prompt cache hit ~98%". ZONE 3 — "Specialised workers": 2×2 hexagons "Researcher" / "Coder" / "Critic" / "Writer", each with glyph + status ribbon ("idle", "running step 3/5", "done", "running step 2/4"). Centre labeled "async message bus". ZONE 4 — "Tools & memory": (a) "Tool registry" panel listing "web_search ×41", "python_exec ×27", "read_file ×18", "write_file ×12", "browser_use ×7"; (b) "Memory" panel with "Short-term scratchpad" and cylinder "Long-term vector store — 1.8M episodes". Bottom inset "Example trace": 8-step horizontal timeline chips from "User asks" through "Planner decomposes", "Researcher: web_search(...)", "Coder: python_exec(...)", "Critic: verify", "Re-plan" (loop-back arrow), "Writer: compose final answer". Title: "Agentic LLM system: planner orchestrates specialised workers over a shared tool and memory layer". Subtitle: "adapted from AutoGen (Wu et al., 2023), LangGraph, and Anthropic Managed Agents patterns".
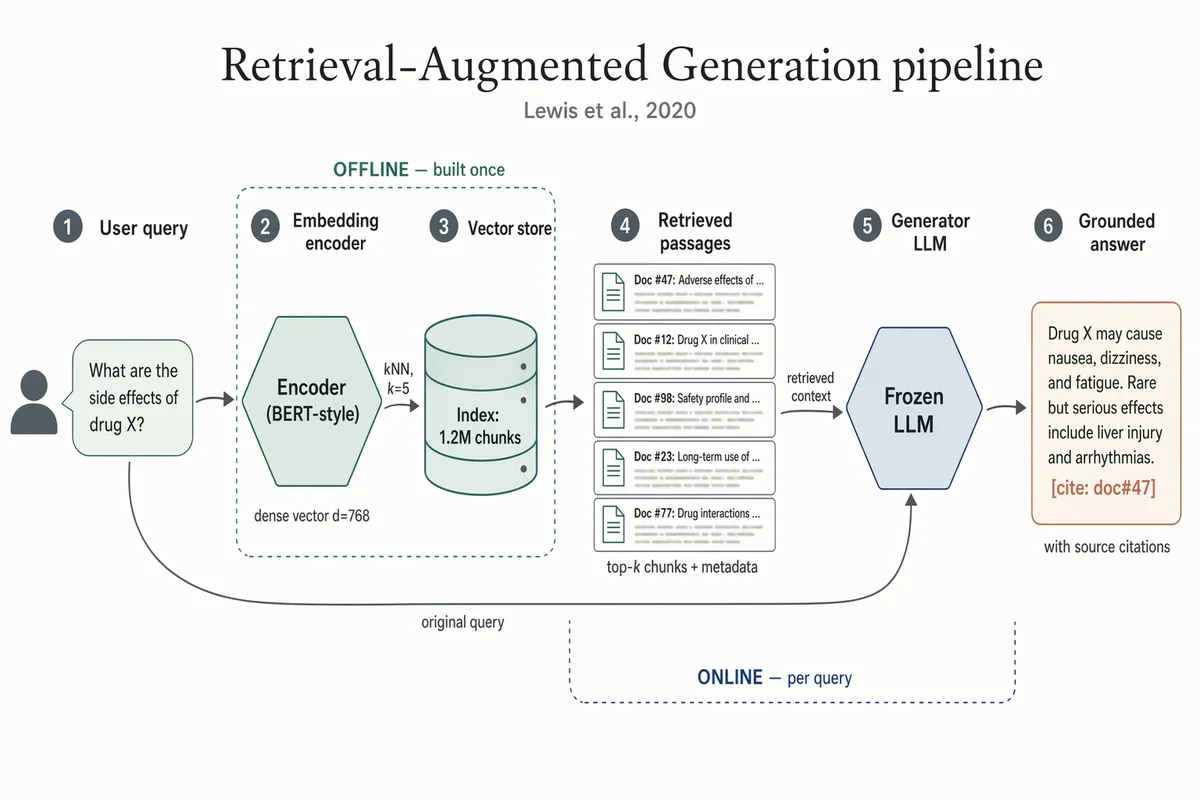
RAG(検索拡張生成) パイプライン
Landscape 16:9 academic systems diagram of a RAG pipeline, 6-stage left-to-right flow. (1) "User query" box with placeholder text "What are the side effects of drug X?" and a small user silhouette. (2) Hexagonal "Embedding encoder (BERT-style)", caption "dense vector d=768". (3) Stylised database cylinder "Vector store" with "Index: 1.2M chunks"; arrow from (2) labeled "kNN, k=5". (4) "Retrieved passages" — stack of 5 doc thumbnails; caption "top-k chunks + metadata". (5) Hexagonal hub "Frozen LLM"; long curved arrow from (1) labeled "original query" also lands here; arrow from (4) labeled "retrieved context". (6) "Grounded answer" with inline marker "[cite: doc#47]"; caption "with source citations". Dashed outline around (2)-(3) labeled "OFFLINE — built once". Dashed outline around (4)-(5) labeled "ONLINE — per query". Title: "Retrieval-Augmented Generation pipeline". Subtitle: "Lewis et al., 2020".

Transformer エンコーダ・デコーダ・アーキテクチャ
Landscape 16:9 academic concept figure of the Transformer encoder-decoder architecture, NeurIPS camera-ready style. Two vertical column stacks side-by-side with a dashed divider. LEFT column header: "ENCODER (×N)". Blocks bottom-to-top: "Input tokens" → "Input Embedding" → "+ Positional Encoding" → dashed "Encoder layer" containing "Multi-Head Self-Attention", "Add & Norm", "Feed-Forward", "Add & Norm", with thin curved residual arrows around each sublayer. RIGHT column header: "DECODER (×N)". Blocks bottom-to-top: "Output tokens (shifted right)" → "Output Embedding" → "+ Positional Encoding" → dashed "Decoder layer" containing "Masked Multi-Head Self-Attention", "Add & Norm", "Multi-Head Cross-Attention" (horizontal arrow from encoder top labeled "keys, values"), "Add & Norm", "Feed-Forward", "Add & Norm". Above decoder: "Linear", "Softmax", "Output probabilities". Title: "Transformer: encoder–decoder with multi-head attention". Subtitle: "Vaswani et al., 2017".
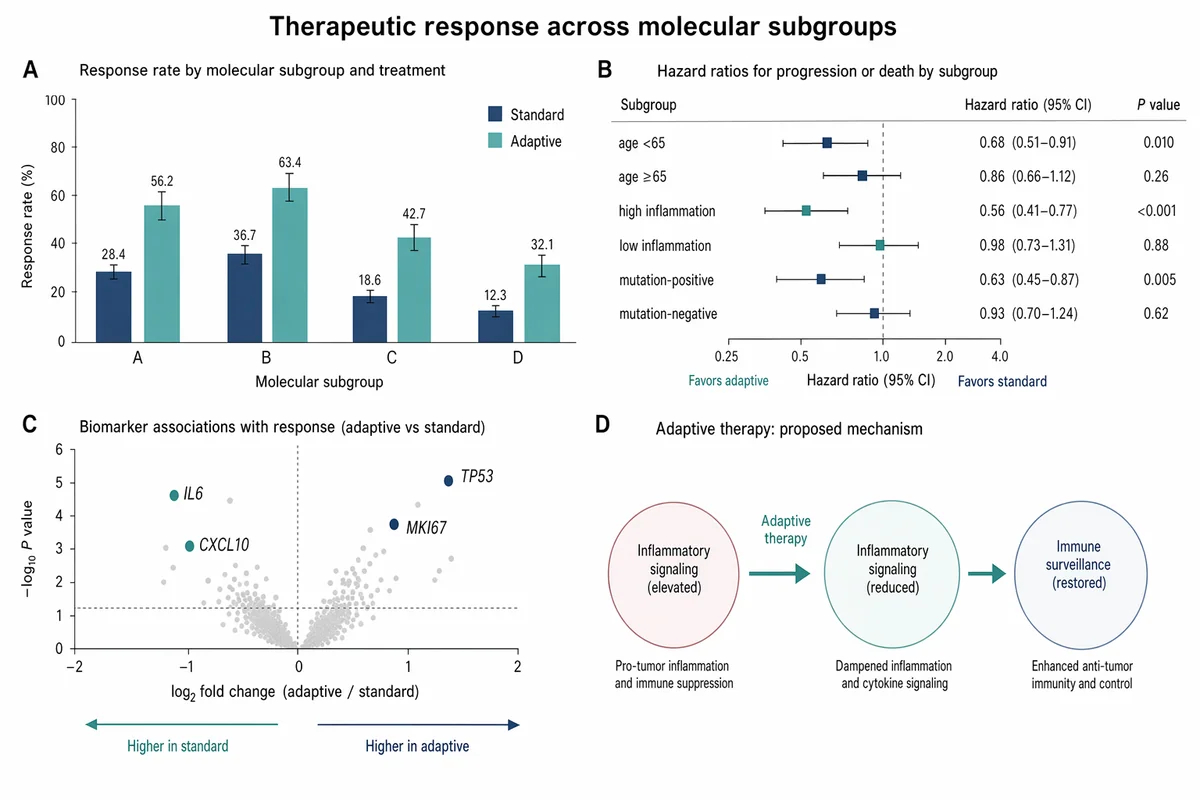
治療応答 バーチャート&フォレストプロット
Create a Nature Medicine style statistical results figure, landscape 3:2 (1536×1024), soft, restrained, publication-quality. Figure title: "Therapeutic response across molecular subgroups". Layout: 4-panel figure labeled A–D. A. Grouped bar chart: response rate (%) for four subgroups "A", "B", "C", "D" across two treatments "standard" and "adaptive". Use muted navy and soft teal bars, thin error bars, numeric labels. B. Forest plot of hazard ratios for subgroups with a vertical reference line at HR=1.0; rows "age <65", "age ≥65", "high inflammation", "low inflammation", "mutation-positive", "mutation-negative". Use small squares and confidence intervals. C. Volcano-style biomarker association plot with pale gray background points and highlighted labeled markers "IL6", "CXCL10", "TP53", "MKI67". D. Minimal mechanism schematic: adaptive therapy reduces inflammatory signaling and restores immune surveillance; use three clean nodes connected by arrows, no complex biology drawings. Style requirements: literature-science aesthetic, white background, soft desaturated colors, thin gray axes, clear legends, compact labels, generous margins, Nature-style figure polish, no fake values that look too random, no decorative background, no watermark.